HistRegGUI
Desktop GUI for histology image registration using DeeperHistReg presets (initial, rigid, nonrigid) designed for practical serial section alignment workflows.
Overview
HistRegGUI provides a simple interface to run common histology registration stages on pairs of images (source → target). It is useful for aligning serial sections, preparing datasets for 3D reconstruction, and supporting downstream quantitative analysis (segmentation, spatial features, etc.).
Key features
- Clear workflow for initial, rigid, and nonrigid registration
- Preset-based execution using DeeperHistReg settings
- Exports registered image(s) and transformation outputs
- Designed to run on CPU-only machines for accessibility
Quick start
- Select moving/source image and fixed/target image.
- Choose a registration preset (initial / rigid / nonrigid).
- Set output folder and run.
Typical output includes the registered image and intermediate results depending on the chosen stage.
Recommended use
- Start with initial when images have large shifts/rotations
- Use rigid for global alignment
- Apply nonrigid for local tissue deformation correction
Inputs & notes
- Works best when images are comparable in scale and staining conditions.
- For large WSI-derived regions, consider working on consistent magnification/patch level.
- Keep file naming consistent to help batch processing and reproducibility.
Troubleshooting
- Output looks misaligned: try running initial first or verify image orientation.
- Too slow: use smaller ROIs/patches or reduce resolution for a first pass.
- Memory issues: avoid extremely large images; pre-crop representative regions.
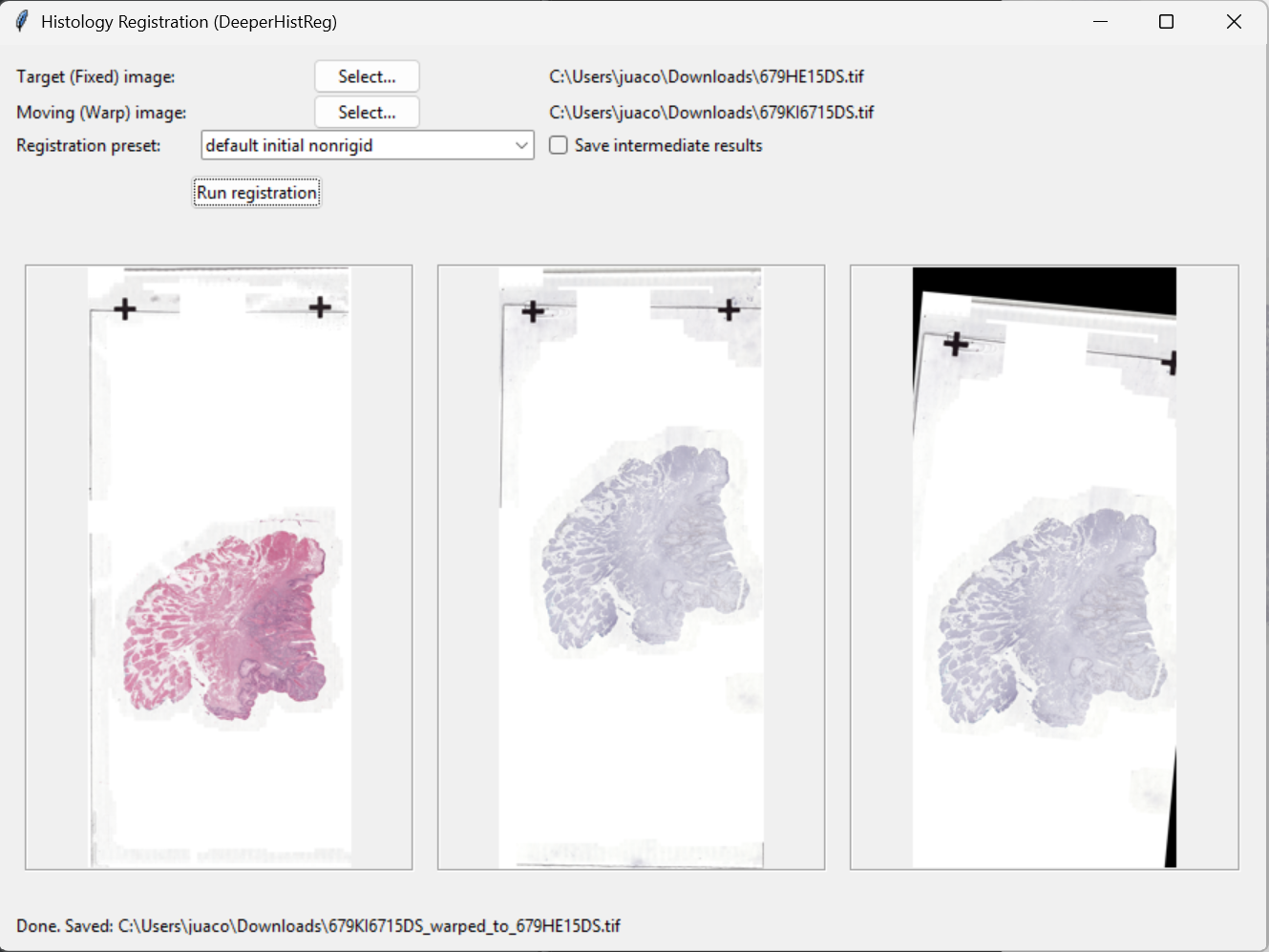
Screenshots
Example interface view.