TiffCropper
Standalone Windows application to extract high-resolution ROIs from large TIFF microscopy images while preserving calibration metadata.
Overview
Designed for digital histopathology researchers who need precise crops from large TIFF files, keeping calibration metadata (resolution, physical size) intact.
Key features
- Browse any .tif/.tiff file
- Coordinate-based ROI input (X, Y, Width, Height)
- Custom suffix for output naming
- Portable executable (no installation)
Quick start
- Click Browse TIFF and select a .tif/.tiff file.
- Enter ROI coordinates: X, Y, Width, Height.
- Set a suffix (optional) and click CROP & SAVE.
Output is saved in the same folder as the original file, using: {original}_crop_{suffix}.tif
Coordinate system
The image is treated as a matrix: X is the column index (left→right), Y is the row index (top→bottom). Width/Height define the ROI size in pixels.
Troubleshooting
- No file selected: ensure the TIFF is not open in ImageJ/Fiji.
- Cannot read dimensions: verify the file is a valid TIFF.
- Access denied: run as Administrator or move the source to a writable folder.
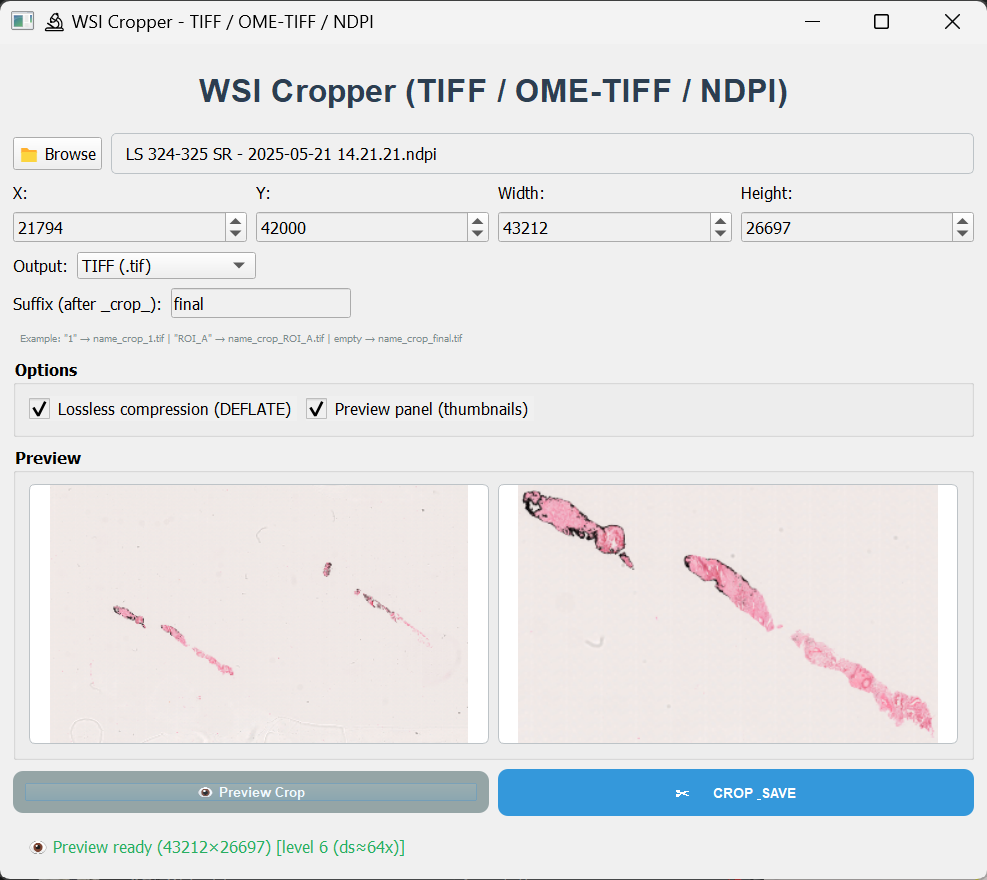
Screenshots
Visualization of the tool.